食品安全是全球公共卫生问题的重要一环,根据世界卫生组织(World Health Organization,WHO)统计,全球约有6亿 人因食用受污染的食物而患病,42万 人死亡[1]。食品会受到一些天然毒素及外源化学危害物的影响,如生物毒素[2]、农药残留、兽药残留、非法食品添加剂和重金属等[3]。人们长期暴露在化学危害物中,会增加罹患中枢神经系统、心血管系统、肾脏、皮肤和生殖系统的急性和慢性疾病以及各种癌症的风险[4]。其中,重金属和环境污染物的危害较高,特别是对儿童、孕妇和免疫力较弱的人群影响更大[5];农药残留和某些加工有害物质的危害程度取决于暴露量和时间;食品添加剂的危害一般较低,但过量或长期摄入仍存在一定风险[6]。目前评估化学危害物对人体健康影响的手段包括血液检查、尿液检测、组织活检和内窥镜检查等[7],这些手段往往是有创性的,需要测定关键生理和组织参数。因此,亟需开发无创且准确的检测方法,以便更有效地进行风险评估,防止食品安全化学危害物带来的严重后果。
近年来,研究表明肠道菌群与人体健康息息相关[8-9]。肠道菌群被视为反映宿主生理变化的一个“微生物器官”,其改变与血糖、血脂和免疫等生理的异常相关[10-12]。例如,肠道中大肠杆菌、Dorea和Peptoniphilus与非酒精性脂肪性肝病风险增加显著相关[13];Akkermansia muciniphila的减少可作为2型糖尿病早期诊断的生物标志物[14];Fusobacterium nucleatum是结直肠癌的关键致病因子和微生物标记物[15]。然而,肠道菌群的结构和组成会受到外源化学物的影响,导致肠道菌群失调[16]。大量的研究表明,抗生素、重金属、持久性有机污染物、杀虫剂和食品添加剂等化学危害物可通过多种机制改变人体肠道菌群的组成和功能(图1),影响肠道稳态[17-20]。例如,诱导肠道菌群失调、损害肠道屏障、引发肠道炎症、增加氧化应激、引起DNA损伤和产生细胞毒性等。有关食品安全化学品对肠道菌群组成和功能影响的关注不断增加,2023年12月联合国粮食及农业组织举办了以“食品安全化学品风险评估中的肠道菌群”为议题的技术会议,会议形成了一个专家共识,即更深入地了解膳食中的化学品如何影响肠道菌群和人类健康将有助于确定可用于化学品风险评估的相关微生物靶标。总之,食源性化学危害物可能通过影响肠道菌群对宿主产生长期、深远的健康影响,利用无创性的肠道菌群检测技术进行食品安全化学危害物风险评估具备巨大的应用潜力。
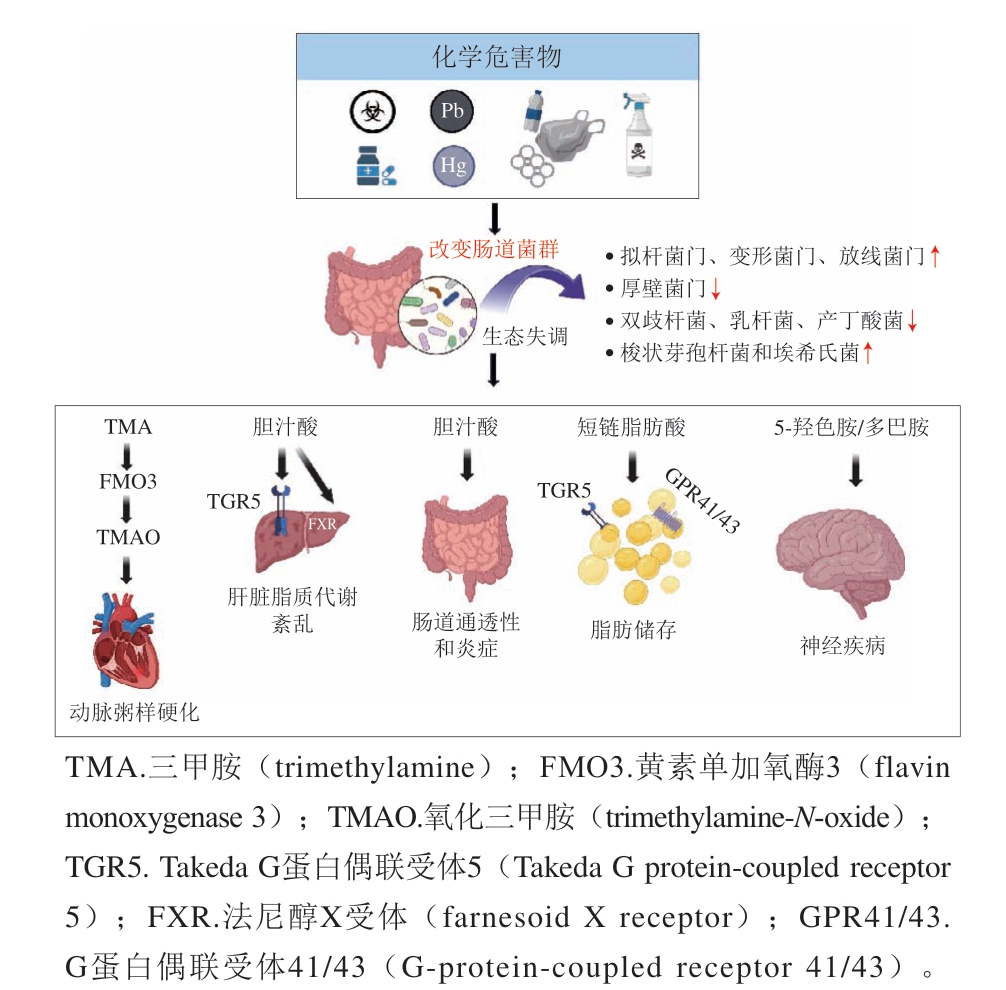
图1 化学危害物对肠道菌群及其对宿主生理的影响
Fig. 1 Effects of chemical hazards on gut microbiota and host physiology
肠道菌群研究的模型或载体种类繁多,包括体外、细胞模型、动物模型和临床试验等。此外,前沿的组学技术如宏基因组学、宏转录组学、代谢组学和宏蛋白质组学也被广泛应用于该领域的研究。这些手段有助于研究人员全面解析食品中化学危害物与肠道菌群的相互作用,为食品安全风险评估工作提供重要参考信息。本综述介绍肠道菌群研究模型和组学技术在食品安全化学危害物风险评估中的应用,以期为食品危害物的调控和管理提供科学依据。同时,为本领域研究人员提供前沿研究进展和相关的方法学工具,为肠道菌群研究模型及组学技术的应用提供理论依据。
1 肠道菌群研究模型在食品安全化学危害物风险评估中的应用
1.1 体外发酵模型
目前,已开发出一系列模拟胃肠道发酵的系统,包括静态发酵模型和动态发酵模型,用于模拟人类对食物或其单一成分的消化以及研究肠道菌群的变化和化学危害物之间的关系[21]。
1.1.1 静态发酵模型
静态发酵是一种简单的体外发酵模型,广泛用于模拟受控条件下肠道菌群与食品安全危害物之间的相互作用。一项体外分批静态培养实验发现,膳食铝暴露显著增加拟杆菌门、变形菌门和放线菌门水平,降低厚壁菌门水平,且改变肠道代谢物水平[22]。此外,一项体外发酵研究表明,膳食乳化剂硬脂酰乳酸钠显著改变肠道菌群,并抑制Roseburia intestinalis和Faecalibacterium prausnitzii等丁酸盐产生者,导致丁酸浓度下降[23]。静态发酵模型简单、经济高效、重现性好、明确实验条件和程序,提高了体外发酵研究的一致性和可比性[24]。然而,也存在一定的局限性,不能反映消化道不同阶段之间的消化转运和阶段性吸收,不能模拟食物消化过程复杂的动态变化以及与宿主生理的相互作用。
1.1.2 动态发酵模型
为了改进静态模型的不足,研究人员开发了一系列的动态发酵模型,通过控制消化过程中的pH值、温度、消化液、酶、肠道蠕动等参数,更真实地模拟肠道中的生理环境和菌群动态变化,探究肠道菌群与宿主相互作用。在所有塑料瓶中,苯二甲酸乙二醇酯(polyethylene terephthalate,PET)是主要的包装材料[25]。Tamargo等[26]为了评估微塑料在消化水平上的潜在风险,通过结合静态模型和动态胃肠道模型(dynamic gastrointestinal simulator,SIMGI),模拟PET在胃肠道中的生物转化过程,发现微塑料喂养会改变人类结肠菌群的组成,对肠腔和黏膜菌群有不同影响[27]。许多体外动态发酵模型显示,食品添加剂如聚山梨醇酯80、三氯蔗糖、二氧化钛、亚硝酸钠和麦芽糊精会改变肠道菌群的组成,减少产丁酸菌,例如Roseburia和粪杆菌,增加与炎症和纤维化相关的细菌,导致短链脂肪酸浓度下降[28-31]。动态发酵系统实验条件控制精确,具有良好的平行性和重现性,长期维持菌群稳定性,可评估食品危害物对菌群影响,且减少动物实验依赖[32]。然而,体外动态发酵系统仍有一定局限性,如难以复现体内复杂生理环境以及菌群与宿主的相互作用[33]。
1.2 体外细胞模型
相较于体外发酵聚焦食品中的危害物与肠道菌群互作,细胞模型能在一定程度上反映出这一过程中宿主的生理活动。单层细胞培养物,特别是癌细胞系或永生细胞系,在胃肠道研究中得到广泛应用[34]。近年来,类器官的三维培养技术得到发展,使得肠道中的隐窝-绒毛单位能够在体外再现,从而更真实地模拟整个肠道微环境。这项技术为研究肠道菌群的生长、代谢及其对宿主的影响提供了更加精确和可靠的模型[35]。
1.2.1 细胞模型
细胞模型是一种通过培养肠道相关细胞(如肠道上皮细胞)构建的体外实验模型,可用于研究肠道细胞与菌群相互作用,探究菌群对肠道屏障功能和免疫调节的影响。Huang Wantang等[36]通过人体Caco-2细胞和肠道菌群的体外模拟实验,探讨了聚乙烯微塑料颗粒(polyethylene microplastics particles,PEMPs)和四溴双酚A(tetrabromobisphenol A,TBBPA)对人体肠道的共同影响,发现二者均恶化Caco-2细胞的状态,高浓度的PEMPs和TBBPA会打破肠道平衡,对人体健康构成威胁。Fournier等[37]通过Caco-2和HT29-MTX肠细胞和肠道菌群的体外共培养实验研究长期暴露于膳食聚乙烯(polyethylene,PE)塑料对婴儿肠道菌群和肠道屏障的影响,发现暴露于PE导致菌群及有益代谢物改变。尽管目前微塑料及其吸附的有机污染物对人体的危害还很小,但随着塑料污染的日益严重,仍需对其潜在的威胁保持警惕。暴露于食品添加剂二氧化钛可损害Caco-2细胞膜完整性,降低线粒体膜电位,并影响紧密连接,破坏肠道屏障[38]。这些发现表明,食品添加剂的安全性评估需要更严格的标准和更广泛的测试。二维细胞培养系统方便,细胞生长迅速,性价比高,但无法模拟天然上皮复杂结构和不同细胞间相互作用,且在重复传代后易产生基因组突变[39]。
1.2.2 类器官模型
由于肠上皮细胞的多样性和复杂性(如吸收细胞、潘氏细胞、杯状细胞、M细胞等),使用单一细胞系无法准确研究所有肠上皮细胞的功能。肠道类器官为肠道表型和功能的模拟提供了重要的突破,能够传代多次而不易基因突变[40]。该模型已被广泛应用于模拟疾病、物质的作用机制和屏障功能的研究[41-43]。最近的一项研究利用小鼠肠道类器官的肠源性单层细胞(enteroid-derived monolayers,EDM)探讨了食品级二氧化钛对肠道的影响,发现暴露于二氧化钛后EDM在细胞增殖、分化、凋亡、遗传毒性、上皮先天防御和紧密连接功能方面的完整性发生改变[44]。肠道类器官模型也被用于食品乳化剂引起肠道屏障损伤的机制分析[45]。研究人员利用M细胞,观察它们在接触微塑料和纳米塑料时发挥的作用,发现高浓度的微粒会显著引发炎症细胞因子的分泌[46]。总体来看,肠道类器官模型在评估食品化学危害物风险方面具有重要的技术应用[47]。三维细胞培养模型能重建肠道组织结构,模拟生化梯度、流动和蠕动[48],但构建和维持类器官模型需使用昂贵的试剂(胶原蛋白基质、生长因子)[49]。此外,由于细菌与细胞共培养会影响细胞的生理特性并造成损伤,因此,很少将细胞与肠道菌群共培养[50]。在研究化学危害物对细胞模型/类器官模型的影响时,无法考虑其对肠道菌群的影响。
1.3 动物模型
如表1所示,小鼠、大鼠、猪和兔子等动物模型常用于研究食品危害物-肠道菌群相互作用和宿主-菌群共生的生物学复杂性。与体外或细胞模型相比,动物模型除了能够良好地模拟人类的体内环境,包括免疫系统、新陈代谢和生理过程之间的相互作用,还能够监测长期暴露下的潜在健康危害。重金属暴露可扰动肠道菌群,进而影响代谢和生理功能,导致宿主损伤。例如,长期接触膳食或者饮用水中镉、铅、铜和铝会以金属特异性和时间依赖性的方式改变无特异性病原体(specific pathogen free,SPF)小鼠的肠道菌群[51-53]。此外,食品添加剂二氧化钛的暴露显著影响SD大鼠的肠道菌群,即使低暴露也会干扰肠道菌群并引起低度肠道炎症[54-55]。小鼠是应用最广泛的动物模型,小鼠肠道菌群与宿主之间的关系能够为理解复杂的危害物-肠道关系提供重要信息[56]。此外,基因编辑小鼠还能够模拟特定疾病状态下的生理和病理特征[57],无菌小鼠模型为菌群功能的因果逻辑分析提供重要支撑[58]。例如,通过将人类粪便菌群移植到无菌小鼠的肠道中,证明了非营养性甜味剂改变的菌群与非葡萄糖不耐受之间的因果和个体化联系[59]。然而,小鼠的肠道生理学与人类有显著差异。小鼠肠道以假长双歧杆菌和动物双歧杆菌为主,而人类则以青春双歧杆菌和长双歧杆菌为主[60]。为此,研究人员在无菌小鼠中移植特定/已知的菌群来建立悉生小鼠[61]。
表1 化学污染物对不同动物模型肠道菌群的影响
Table 1 Effects of chemical pollutants on gut microbiota in different animal models
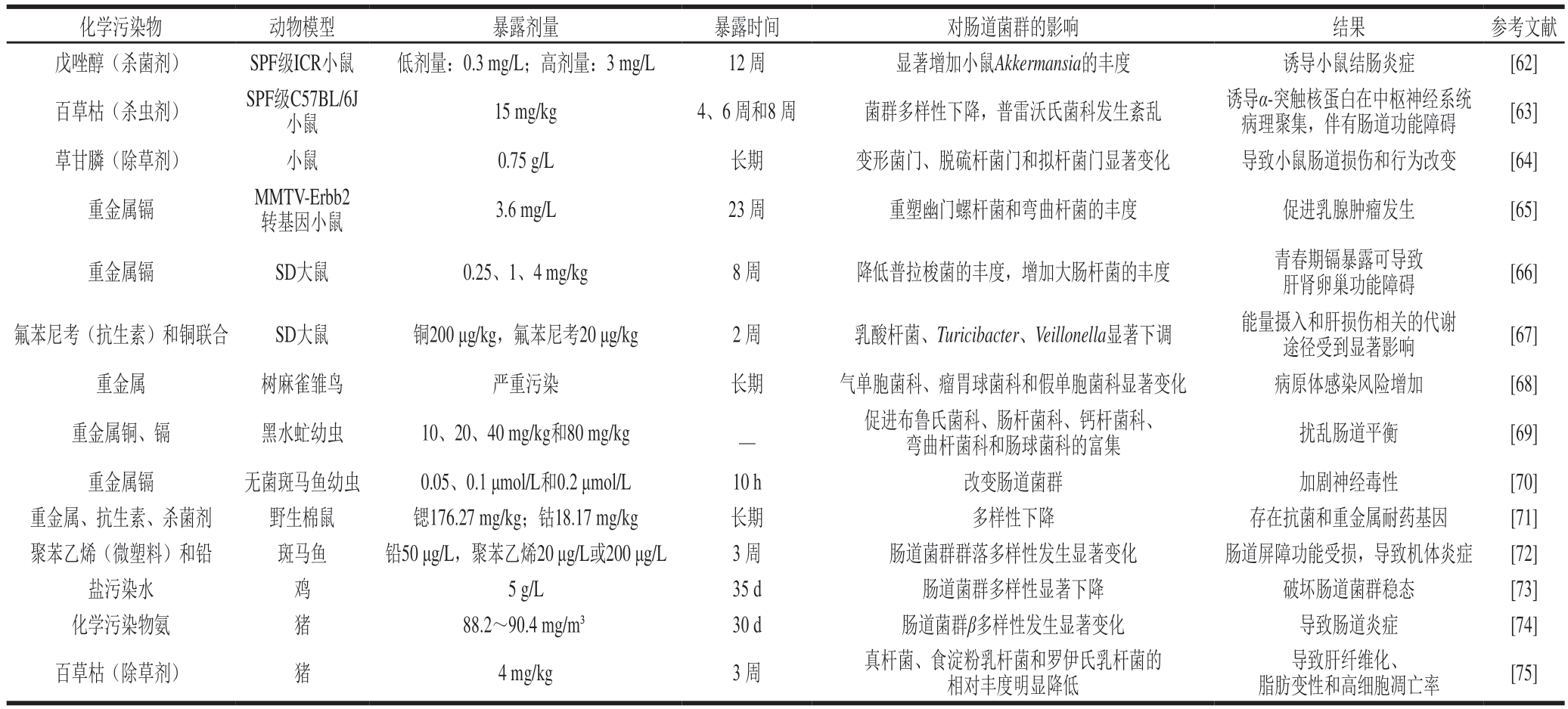
注:—.文献未说明。
化学污染物动物模型暴露剂量暴露时间对肠道菌群的影响结果参考文献戊唑醇(杀菌剂)SPF级ICR小鼠低剂量:0.3 mg/L;高剂量:3 mg/L12 周显著增加小鼠Akkermansia的丰度诱导小鼠结肠炎症[62]百草枯(杀虫剂)SPF级C57BL/6J小鼠15 mg/kg4、6 周和8 周菌群多样性下降,普雷沃氏菌科发生紊乱诱导α-突触核蛋白在中枢神经系统病理聚集,伴有肠道功能障碍[63]草甘膦(除草剂)小鼠0.75 g/L长期变形菌门、脱硫杆菌门和拟杆菌门显著变化导致小鼠肠道损伤和行为改变[64]重金属镉MMTV-Erbb2转基因小鼠3.6 mg/L23 周重塑幽门螺杆菌和弯曲杆菌的丰度促进乳腺肿瘤发生[65]重金属镉SD大鼠0.25、1、4 mg/kg8 周降低普拉梭菌的丰度,增加大肠杆菌的丰度青春期镉暴露可导致肝肾卵巢功能障碍[66]氟苯尼考(抗生素)和铜联合SD大鼠铜200 μg/kg,氟苯尼考20 μg/kg2 周乳酸杆菌、Turicibacter、Veillonella显著下调能量摄入和肝损伤相关的代谢途径受到显著影响[67]重金属树麻雀雏鸟严重污染长期气单胞菌科、瘤胃球菌科和假单胞菌科显著变化病原体感染风险增加[68]重金属铜、镉黑水虻幼虫10、20、40 mg/kg和80 mg/kg_促进布鲁氏菌科、肠杆菌科、钙杆菌科、弯曲杆菌科和肠球菌科的富集扰乱肠道平衡[69]重金属镉无菌斑马鱼幼虫0.05、0.1 μmol/L和0.2 μmol/L10 h改变肠道菌群加剧神经毒性[70]重金属、抗生素、杀菌剂野生棉鼠锶176.27 mg/kg;钴18.17 mg/kg长期多样性下降存在抗菌和重金属耐药基因[71]聚苯乙烯(微塑料)和铅斑马鱼铅50 μg/L,聚苯乙烯20 μg/L或200 μg/L3 周肠道菌群群落多样性发生显著变化肠道屏障功能受损,导致机体炎症[72]盐污染水鸡5 g/L35 d肠道菌群多样性显著下降破坏肠道菌群稳态[73]化学污染物氨猪88.2~90.4 mg/m³30 d肠道菌群β多样性发生显著变化导致肠道炎症[74]百草枯(除草剂)猪4 mg/kg3 周真杆菌、食淀粉乳杆菌和罗伊氏乳杆菌的相对丰度明显降低导致肝纤维化、脂肪变性和高细胞凋亡率[75]
1.4 临床试验
临床研究可直接观察食品安全危害物对人类肠道菌群的影响。队列研究表明,产前接触食品增塑剂(如多溴二苯醚和多氯联苯)会导致胎儿肠道菌群变异,增加拟杆菌属、丙酸杆菌属和丙酸杆菌科的相对丰度[76]。这表明化学暴露的影响可能具有长期性,甚至延续到后代的肠道健康。一项成人的横断面研究表明,铅暴露会改变肠道菌群的组成,增加肠道菌群α多样性,提高变形菌门的丰度[77]。膳食乳化剂羧甲基纤维素的增加导致志愿者餐后腹部不适,改变肠道菌群的组成,增加菌群对正常无菌内黏液层的侵占,导致肠道炎症[78]。非营养性甜味剂(糖精、三氯蔗糖、阿斯巴甜和甜叶菊)干预均显著改变粪便菌群以及血浆代谢组,且糖精和三氯蔗糖显著损害血糖反应[59]。然而,这些结果仅基于短期观察,长时间暴露以及暴露剂量的影响仍需进一步探索。临床试验的优点在于可以通过直接观察提供食品安全化学品对人体肠道菌群影响的最直接相关数据,且能涵盖不同年龄、国籍和地理位置的多样性和代表性。当然,临床试验也存在诸多挑战,包括难以控制混杂因素、人员失访、数据分析复杂、成本高昂以及涉及资源和伦理问题[79]。
2 肠道菌群研究组学技术在食品安全化学危害物风险评估中的应用
目前的研究方法中,宏基因组测序技术能够在种水平甚至株水平展示肠道菌群的组成、结构及功能[80]。然而,微生物的存在并不能表明其在肠道中的作用,功能基因的存在也无法反映基因表达情况。宏转录组学提供了在RNA水平上的基因表达数据[81],宏蛋白组学则提供了在功能实现最终阶段的蛋白质数据,直接反映细胞的功能状态。三者相互补充,提供从基因到蛋白质表达的全过程视图,有助于全面地理解生物体的生理状态和功能。代谢组学则有助于识别菌群代谢物及宿主-菌群共代谢物,揭示宿主与肠道菌群间的复杂相互作用[82]。宏基因组学、宏蛋白组学、宏转录组学、代谢组学和培养组学通过结合宿主和微生物的多元数据揭示宿主遗传与微生物互作,为研究食品安全危害物对菌群功能的影响提供了新思路。
组学技术可以揭示饮食中暴露有毒化合物后引发疾病、肠道紊乱和生态失调的因果关系及分子机制[83]。探索菌群和代谢物之间的变化和关系对于改善包括重金属中毒在内的疾病的预防、诊断和治疗至关重要。代谢组学与宏基因组学的结合,可用于检测铅对肠道菌群的毒性及其对人类健康的影响[84],发现铅暴露改变了鞘脂代谢和离子转运水平[85]。这可能为揭示铅毒性机制提供了一种新方法,但是未来需要更多的研究证实研究结果的普适性。大米是膳食中镉摄入量的最大来源,亚急性镉暴露可破坏肠道微生态平衡和甘油磷脂代谢稳态。宏基因组和代谢组学的结合揭示了亚急性镉暴露大鼠与健康大鼠在菌群及甘油磷脂代谢上的差异,为生物体镉中毒的预测、预警和靶向治疗提供了新方向[66,86]。汞暴露会造成小龙虾组织损伤,扰乱氧化还原平衡,破坏离子稳态,降低免疫反应,导致细胞凋亡,降低细菌多样性,促进肠道菌群失调[87]。该研究从组织、细胞、分子和微生物层面阐明了汞对水生甲壳类动物的毒性机制。除草剂草甘膦暴露会抑制胃肠道菌群的莽草酸途径,降低多种代谢物水平,引起氧化应激反应[88]。这些数据表明,肠道菌群及其代谢物的变化或许可作为食品安全危害物暴露的敏感指标。此外,粪便宏转录组可揭示暴露化学危害物后代谢途径的变化,阐明菌群结构变化如何影响人类健康的分子机制。例如,Defois等[89]借助于宏转录组学手段发现,食源性化学污染物(多环芳烃、多氯联苯、溴化阻燃剂、二噁英、农药和杂环胺)会导致与核糖体、翻译和核酸结合相关的微生物转录水平下降,促进脂质代谢过程以及质膜、周质空间、蛋白激酶活性和受体活性相关的微生物基因表达增加,通过干扰肠道菌群代谢物促进肠道炎症状态的建立。
多组学技术(宏基因组学、宏蛋白组学和宏转录组学等)在研究食品中化学危害物对肠道菌群及宿主的影响方面,具有综合分析、动态跟踪和揭示机制等优势,能够提供高分辨率的全面数据,揭示个体差异和生物学网络的复杂相互作用。例如,随着测序深度的增加,宏基因组学可用于物种和亚种水平的鉴定,结合转录组学还可以深入分析功能及差异基因表达,有助于了解在危害物暴露下肠道菌群中活跃成员、基因和途径的变化[90]。然而,随着测序深度的增加,对应的成本更高[91]。测序生成的大量数据需要高性能计算工具进行分析和存储,需要掌握专业的技术生物信息学和科学专业知识才能分析。
3 存在的挑战
化学危害物和肠道菌群之间存在一定的相关性,危害物可直接影响肠道菌群,改变其代谢产物和免疫系统,进而对宿主健康构成潜在风险。反过来,肠道菌群也能够代谢和转化某些化学物质,还能通过调节宿主的免疫系统、屏障功能和代谢途径影响化学物质的毒性和生物利用度。使用肠道菌群作为食品安全化学危害物风险评估的新靶标具有巨大潜力,但也面临诸多挑战。
3.1 难以确定危害物对菌群的效应
肠道菌群的组成在整个成年期保持相对稳定[92],但可能受到宿主遗传、暴露组、饮食、年龄、出生方式和抗生素等因素的影响[93]。然而,这些因素对肠道菌群的影响程度和机制尚不清楚[94],由于众多难以控制的混杂因素,在解释化学危害物对肠道菌群变异时,确定其效应大小具有一定困难。为了消除菌群相关干扰变量对最终实验结果的影响,可采用线性混合模型框架中的统计调整、配对设计、均一化实验条件,以及随机化和扩大样本量等策略,尽量减少这些因素的干扰[95-96]。此外,因果关系通常也是困扰研究人员的难题之一,肠道菌群与宿主之间的相互作用复杂,是生理的改变引起菌群的改变还是菌群的改变导致生理的异常,确定其因果关系需要深入的机制研究和严谨的实验设计。
3.2 数据可访问性差
组学数据通常存储在NCBI SRA、EBI SRA、HMP以及NGDC等不同的平台中,但其存储格式各异,导致数据统一访问存在困难。许多数据库要求研究者进行注册和数据请求流程,进一步增加了获取数据的难度。此外,大规模组学数据集的下载和处理对计算资源和存储空间的需求极高,导致数据获取过程复杂且耗时。
3.3 跨组学数据整合不佳
目前的数据集和数据库远远不足以构建有效的预测模型,以准确理解污染物与肠道菌群之间的关系。虽然很多研究团队分别进行了暴露组学[97-98]和肠道菌群研究[99-101],但这些数据难以有效整合。其次,不同类型的组学数据格式和标准不一致。例如,宏基因组侧重于基因层面的信息,宏蛋白组则侧重于蛋白质层面的表征,而转录组数据则关注基因表达水平,这些不同层面的数据在整合时面临挑战。为了应对这些挑战,研究人员通常借助降维和聚类、网络、相关性、空间多组学和代谢建模等方法优化跨组学数据整合的算法。
3.4 数据质量要求高
粪便细菌群落的检测会受到各种因素的影响,包括样品采集和测序。例如,粪便样本采集和保存[102]、微生物基因组DNA提取方法[103]、测序文库制备策略、测序平台选择[104]、测序深度[105]、细菌数据库选择和生物信息学流程[106],这些步骤均可能影响最终测序数据的精度和可靠性。多年来,人们已经为宏基因组学的标准化做出了多项努力,包括开发微生物参考标准[107]、考虑最佳实践[108]以及DNA提取的标准方法[109]。最终,形成了一套经过验证的方案和方法指导,以提高基于宏基因组学的人类粪便菌群分析的准确性和重现性[110]。
3.5 无共识的数据分析思路
宏基因组数据分析目前尚无统一的共识方法,数据分析思路呈现出多样化的特点。由于数据的复杂性和多维度特征,不同研究团队采用的分析思路和方法各异,涉及物种注释、功能注释和宏基因组序列组装等。这种多样化处理导致了数据解读的困难和结果的可比性问题。未来可以建立标准化的数据分析指南,开发和推广标准化分析工具和平台,促进数据共享和跨学科合作。
3.6 缺乏“金标准”方法
每种组学技术各有其独特的优势与局限,将宏基因组学、代谢组学以及转录组学全面应用于每一个样本存在较高的时间和经济成本,通常产生海量难以分析和解释的数据,面临着高级生物信息学数据分析的技术瓶颈。此外,虽然许多研究模型可以确定化学危害物和肠道菌群的相互作用,如体外、细胞及动物模型可控、可重复,但无法完成模拟人体复杂生理环境,将实验结果转化为临床应用具有一定的挑战性。许多肠道细菌菌株在肠道中的丰度很低或难以培养,制约了其在体外的应用[111]。总之,选择组学技术或模型手段没有统一的“金标准”方法,需要结合具体的研究目标和样本特性选择适当的技术和方法。
4 结 语
食品安全化学危害物对肠道菌群组成、代谢功能及肠道稳态具有多重影响,并与多种疾病的发生发展有关。因此,肠道菌群结构和功能的变化在食品安全化学危害物风险评估中具有重要意义。肠道菌群研究模型和组学技术从不同层面揭示了化学危害物和菌群相互作用的动态过程,提供了了解危害物在基因表达、蛋白质合成和代谢调控中的作用的新视角。然而,将肠道菌群作为风险评估靶标的实际应用仍有待发展。未来可以利用人工智能和机器学习的手段进行组学数据的分析和解释,结合生物信息学资源,建立食品不良反应数据库,从而更快、更准确地分析大型数据集,为相应的食品安全性风险评估提供参考信息。
[1] ZHAO Y L, BHAVYA M L, PATANGE A, et al. Plasma-activated liquids for mitigating biofilms on food and food contact surfaces[J].Comprehensive Reviews in Food Science and Food Safety, 2023,22(3): 1654-1685. DOI:10.1111/1541-4337.13126.
[2] TANG X Q, ZUO J S, YANG C, et al. Current trends in biosensors for biotoxins (mycotoxins, marine toxins, and bacterial food toxins):principles, application, and perspective[J]. TrAC Trends in Analytical Chemistry, 2023, 165: 117144. DOI:10.1016/j.trac.2023.117144.
[3] ONG K J, JOHNSTON J, DATAR I, et al. Food safety considerations and research priorities for the cultured meat and seafood industry[J].Comprehensive Reviews in Food Science and Food Safety, 2021,20(6): 5421-5448. DOI:10.1111/1541-4337.12853.
[4] NAIDU R, BISWAS B, WILLETT I R, et al. Chemical pollution:a growing peril and potential catastrophic risk to humanity[J].Environment International, 2021, 156: 106616. DOI:10.1016/j.envint.2021.106616.
[5] ZHENG K Y, ZENG Z J, TIAN Q W, et al. Epidemiological evidence for the effect of environmental heavy metal exposure on the immune system in children[J]. Science of the Total Environment, 2023, 868:161691. DOI:10.1016/j.scitotenv.2023.161691.
[6] WU L, ZHANG C H, LONG Y X, et al. Food additives: from functions to analytical methods[J]. Critical Reviews in Food Science and Nutrition, 2022, 62(30): 8497-8517. DOI:10.1080/10408398.2021.1929823.
[7] DE SANTIAGO E R, DINIS-RIBEIRO M, POHL H, et al. Reducing the environmental footprint of gastrointestinal endoscopy: European Society of Gastrointestinal Endoscopy (ESGE) and European Society of Gastroenterology and Endoscopy Nurses and Associates (ESGENA)position statement[J]. Endoscopy, 2022, 54(8): 797-826. DOI:10.1055/a-1859-3726.
[8] YANG J P, YANG X L, WU G J, et al. Gut microbiota modulate distal symmetric polyneuropathy in patients with diabetes[J]. Cell Metabolism,2023, 35(9): 1548-1562.e7. DOI:10.1016/j.cmet.2023.06.010.
[9] LIU Y, WANG Y, NI Y Q, et al. Gut microbiome fermentation determines the efficacy of exercise for diabetes prevention[J]. Cell Metabolism, 2020, 31(1): 77-91.e5. DOI:10.1016/j.cmet.2019.11.001.
[10] MULLER P A, MATHEIS F, SCHNEEBERGER M, et al. Microbiotamodulated CART+ enteric neurons autonomously regulate blood glucose[J]. Science, 2020, 370: 314-321. DOI:10.1126/science.abd6176.
[11] WEN J J, LI M Z, CHEN C H, et al. Tea polyphenol and epigallocatechin gallate ameliorate hyperlipidemia via regulating liver metabolism and remodeling gut microbiota[J]. Food Chemistry, 2023,404: 134591. DOI:10.1016/j.foodchem.2022.134591.
[12] FIDELLE M, RAUBER C, SILVA C A C, et al. A microbiotamodulated checkpoint directs immunosuppressive intestinal T cells into cancers[J]. Science, 2023, 380: eabo2296. DOI:10.1126/science.abo2296.
[13] ARON-WISNEWSKY J, VIGLIOTTI C, WITJES J, et al. Gut microbiota and human NAFLD: disentangling microbial signatures from metabolic disorders[J]. Nature Reviews Gastroenterology &Hepatology, 2020, 17(5): 279-297. DOI:10.1038/s41575-020-0269-9.
[14] ADESHIRLARIJANEY A, GEWIRTZ A T. Considering gut microbiota in treatment of type 2 diabetes mellitus[J]. Gut Microbes,2020, 11(3): 253-264. DOI:10.1080/19490976.2020.1717719.
[15] WANG N, FANG J Y. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer[J]. Trends in Microbiology, 2023, 31(2): 159-172. DOI:10.1016/j.tim.2022.08.010.
[16] LI X Q, NIU H X, HUANG Z L, et al. Deciphering the role of the gut microbiota in exposure to emerging contaminants and diabetes: a review[J]. Metabolites, 2024, 14(2): 108. DOI:10.3390/metabo14020108.
[17] ELMASSRY M M, ZAYED A, FARAG M A. Gut homeostasis and microbiota under attack: impact of the different types of food contaminants on gut health[J]. Critical Reviews in Food Science and Nutrition, 2022, 62(3): 738-763. DOI:10.1080/10408398.2020.1828263.
[18] LI X, TAN C P, LIU Y F, et al. Interactions between food hazards and intestinal barrier: impact on foodborne diseases[J]. Journal of Agricultural and Food Chemistry, 2020, 68(50): 14728-14738.DOI:10.1021/acs.jafc.0c07378.
[19] WANG C, YUAN Z X, LI J J, et al. Acute effects of antimony exposure on adult zebrafish (Danio rerio): from an oxidative stress and intestinal microbiota perspective[J]. Fish & Shellfish Immunology, 2022, 123:1-9. DOI:10.1016/j.fsi.2022.02.050.
[20] BAO L J, WEI Y L, YAO Y, et al. Global trends of research on emerging contaminants in the environment and humans: a literature assimilation[J].Environmental Science and Pollution Research International, 2015, 22(3):1635-1643. DOI:10.1007/s11356-014-3404-8.
[21] KOUTSOS A, LIMA M, CONTERNO L, et al. Effects of commercial apple varieties on human gut microbiota composition and metabolic output using an in vitro colonic model[J]. Nutrients, 2017, 9(6): 533.DOI:10.3390/nu9060533.
[22] YU L L, DUAN H, KELLINGRAY L, et al. Lactobacillus plantarummediated regulation of dietary aluminum induces changes in the human gut microbiota: an in vitro colonic fermentation study[J]. Probiotics and Antimicrobial Proteins, 2021, 13(2): 398-412. DOI:10.1007/s12602-020-09677-0.
[23] ELMÉN L, ZLAMAL J E, SCOTT D A, et al. Dietary emulsifier sodium stearoyl lactylate alters gut microbiota in vitro and inhibits bacterial butyrate producers[J]. Frontiers in Microbiology, 2020, 11:892. DOI:10.3389/fmicb.2020.00892.
[24] EGGER L, MÉNARD O, DELGADO-ANDRADE C, et al. The harmonized INFOGEST in vitro digestion method: from knowledge to action[J]. Food Research International, 2016, 88: 217-225.DOI:10.1016/j.foodres.2015.12.006.
[25] BECERRIL-ARREOLA R, BUCKLIN R E. Beverage bottle capacity,packaging efficiency, and the potential for plastic waste reduction[J].Scientific Reports, 2021, 11(1): 3542. DOI:10.1038/s41598-021-82983-x.
[26] TAMARGO A, MOLINERO N, REINOSA J J, et al. PET microplastics affect human gut microbiota communities during simulated gastrointestinal digestion, first evidence of plausible polymer biodegradation during human digestion[J]. Scientific Reports, 2022,12(1): 528. DOI:10.1038/s41598-021-04489-w.
[27] YAN Z H, ZHANG S H, ZHAO Y G, et al. Phthalates released from microplastics inhibit microbial metabolic activity and induce different effects on intestinal luminal and mucosal microbiota[J]. Environmental Pollution, 2022, 310: 119884. DOI:10.1016/j.envpol.2022.119884.
[28] GONZA I, GOYA-JORGE E, DOUNY C, et al. Food additives impair gut microbiota from healthy individuals and IBD patients in a colonic in vitro fermentation model[J]. Food Research International, 2024,182: 114157. DOI:10.1016/j.foodres.2024.114157.
[29] GERASIMIDIS K, BRYDEN K, CHEN X F, et al. The impact of food additives, artificial sweeteners and domestic hygiene products on the human gut microbiome and its fibre fermentation capacity[J].European Journal of Nutrition, 2020, 59(7): 3213-3230. DOI:10.1007/s00394-019-02161-8.
[30] VAN DEN ABBEELE P, POPPE J, DEYAERT S, et al. Low-no-calorie sweeteners exert marked compound-specific impact on the human gut microbiota ex vivo[J]. International Journal of Food Sciences and Nutrition,2023, 74(5): 630-644. DOI:10.1080/09637486.2023.2240037.
[31] CASTAÑEDA-MONSALVE V, FRÖHLICH L F, HAANGE S B, et al.High-throughput screening of the effects of 90 xenobiotics on the simplified human gut microbiota model (SIHUMIx): a metaproteomic and metabolomic study[J]. Frontiers in Microbiology, 2024, 15:1349367. DOI:10.3389/fmicb.2024.1349367.
[32] MA F M, LUO Y S, LIU Y Q, et al. The disruption on gut microbiome of decabromodiphenyl ethane exposure in the simulator of the human intestinal microbial ecosystem (SHIME)[J]. Toxicology and Applied Pharmacology, 2022, 452: 116194. DOI:10.1016/j.taap.2022.116194.
[33] CHEN P, SUN J W, LIANG Z Q, et al. The bioavailability of soy isoflavones in vitro and their effects on gut microbiota in the simulator of the human intestinal microbial ecosystem[J].Food Research International, 2022, 152: 110868. DOI:10.1016/j.foodres.2021.110868.
[34] LAU H C H, KRANENBURG O, XIAO H P, et al. Organoid models of gastrointestinal cancers in basic and translational research[J]. Nature Reviews Gastroenterology & Hepatology, 2020, 17(4): 203-222.DOI:10.1038/s41575-019-0255-2.
[35] XIANG Y Q, WEN H, YU Y, et al. Gut-on-chip: recreating human intestine in vitro[J]. Journal of Tissue Engineering, 2020, 11:2041731420965318. DOI:10.1177/2041731420965318.
[36] HUANG W T, YIN H, YANG Y Y, et al. Influence of the coexposure of microplastics and tetrabromobisphenol A on human gut:simulation in vitro with human cell Caco-2 and gut microbiota[J].Science of the Total Environment, 2021, 778: 146264. DOI:10.1016/j.scitotenv.2021.146264.
[37] FOURNIER E, RATEL J, DENIS S, et al. Exposure to polyethylene microplastics alters immature gut microbiome in an infant in vitro gut model[J]. Journal of Hazardous Materials, 2023, 443: 130383.DOI:10.1016/j.jhazmat.2022.130383.
[38] MARTIN N, WASSMUR B, BAUN A, et al. Availability and effects of n-TiO2 and PCB77 in fish in vitro models of the intestinal barrier and liver under single- and/or co-exposure scenarios[J]. Aquatic Toxicology, 2022, 253: 106343. DOI:10.1016/j.aquatox.2022.106343.
[39] YIN Y B, GUO S G, WAN D, et al. Enteroids: promising in vitro models for studies of intestinal physiology and nutrition in farm animals[J]. Journal of Agricultural and Food Chemistry, 2019, 67(9):2421-2428. DOI:10.1021/acs.jafc.8b06908.
[40] YIN Y B, ZHOU D G. Organoid and enteroid modeling of Salmonella infection[J]. Frontiers in Cellular and Infection Microbiology, 2018, 8:102. DOI:10.3389/fcimb.2018.00102.
[41] BAO L, CUI X J, BAI R, et al. Advancing intestinal organoid technology to decipher nano-intestine interactions and treat intestinal disease[J]. Nano Research, 2023, 16(3): 3976-3990. DOI:10.1007/s12274-022-5150-4.
[42] INABA A, ARINAGA A, TANAKA K, et al. Interleukin-4 promotes tuft cell differentiation and acetylcholine production in intestinal organoids of non-human primate[J]. International Journal of Molecular Sciences, 2021, 22(15): 7921. DOI:10.3390/ijms22157921.
[43] CHENG Y, HALL T R, XU X, et al. Targeting uPA-uPAR interaction to improve intestinal epithelial barrier integrity in inflammatory bowel disease[J]. EBioMedicine, 2022, 75: 103758. DOI:10.1016/j.ebiom.2021.103758.
[44] MALAISÉ Y, CASALE E, PETTES-DULER A, et al. Validating enteroid-derived monolayers from murine gut organoids for toxicological testing of inorganic particles: proof-of-concept with food-grade titanium dioxide[J]. International Journal of Molecular Sciences, 2024, 25(5): 2635. DOI:10.3390/ijms25052635.
[45] OGULUR I, YAZICI D, PAT Y, et al. Mechanisms of gut epithelial barrier impairment caused by food emulsifiers polysorbate 20 and polysorbate 80[J]. Allergy, 2023, 78(9): 2441-2455. DOI:10.1111/all.15825.
[46] CHEN Y, WILLIAMS A M, GORDON E B, et al. Biological effects of polystyrene micro- and nano-plastics on human intestinal organoid-derived epithelial tissue models without and with M cells[J].Nanomedicine: Nanotechnology, Biology and Medicine, 2023, 50:102680. DOI:10.1016/j.nano.2023.102680.
[47] WANG B, IGLESIAS-LEDON L, BISHOP M, et al. Impact of microand nano-plastics on human intestinal organoid-derived epithelium[J].Current Protocols, 2024, 4(4): e1027. DOI:10.1002/cpz1.1027.
[48] TAEBNIA N, RÖMLING U, LAUSCHKE V M. In vitro and ex vivo modeling of enteric bacterial infections[J]. Gut Microbes, 2023, 15(1):2158034. DOI:10.1080/19490976.2022.2158034.
[49] ALMEQDADI M, MANA M D, ROPER J, et al. Gut organoids:mini-tissues in culture to study intestinal physiology and disease[J].American Journal of Physiology: Cell Physiology, 2019, 317(3):C405-C419. DOI:10.1152/ajpcell.00300.2017.
[50] LEE K W, SHIN J S, LEE C M, et al. Gut-on-a-chip for the analysis of bacteria-bacteria interactions in gut microbial community: what would be needed for bacterial co-culture study to explore the diet-microbiota relationship?[J]. Nutrients, 2023, 15(5): 1131. DOI:10.3390/nu15051131.
[51] ZHAI Q X, LI T Q, YU L L, et al. Effects of subchronic oral toxic metal exposure on the intestinal microbiota of mice[J]. Science Bulletin, 2017, 62(12): 831-840. DOI:10.1016/j.scib.2017.01.031.
[52] LIU Y H, LI Y H, XIA Y H, et al. The dysbiosis of gut microbiota caused by low-dose cadmium aggravate the injury of mice liver through increasing intestinal permeability[J]. Microorganisms, 2020,8(2): 211. DOI:10.3390/microorganisms8020211.
[53] WANG J T, HU W, YANG H L, et al. Arsenic concentrations, diversity and co-occurrence patterns of bacterial and fungal communities in the feces of mice under sub-chronic arsenic exposure through food[J]. Environment International, 2020, 138: 105600. DOI:10.1016/j.envint.2020.105600.
[54] CHEN Z J, ZHOU D, HAN S, et al. Hepatotoxicity and the role of the gut-liver axis in rats after oral administration of titanium dioxide nanoparticles[J]. Particle and Fibre Toxicology, 2019, 16(1): 48.DOI:10.1186/s12989-019-0332-2.
[55] MAO Z L, LI Y Q, DONG T Y, et al. Exposure to titanium dioxide nanoparticles during pregnancy changed maternal gut microbiota and increased blood glucose of rat[J]. Nanoscale Research Letters, 2019,14(1): 26. DOI:10.1186/s11671-018-2834-5.
[56] GUO J G, SONG C C, LIU Y B, et al. Characteristics of gut microbiota in representative mice strains: implications for biological research[J].Animal Models and Experimental Medicine, 2022, 5(4): 337-349.DOI:10.1002/ame2.12257.
[57] LI H Y, YANG Y, HONG W Q, et al. Applications of genome editing technology in the targeted therapy of human diseases: mechanisms,advances and prospects[J]. Signal Transduction and Targeted Therapy,2020, 5(1): 1. DOI:10.1038/s41392-019-0089-y.
[58] RIDAURA V K, FAITH J J, REY F E, et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice[J]. Science,2013, 341: 1241214. DOI:10.1126/science.1241214.
[59] SUEZ J, COHEN Y, VALDÉS-MAS R, et al. Personalized microbiome-driven effects of non-nutritive sweeteners on human glucose tolerance[J]. Cell, 2022, 185(18): 3307-3328.e19.DOI:10.1016/j.cell.2022.07.016.
[60] XIAO Y, ZHAO J X, ZHANG H, et al. Colonized niche, evolution and function signatures of Bifidobacterium pseudolongum within bifidobacterial genus[J]. Foods, 2021, 10(10): 2284. DOI:10.3390/foods10102284.
[61] CHOO J M, ROGERS G B. Establishment of murine gut microbiota in gnotobiotic mice[J]. iScience, 2021, 24(2): 102049. DOI:10.1016/j.isci.2021.102049.
[62] MENG Z Y, SUN W, LIU W, et al. A common fungicide tebuconazole promotes colitis in mice via regulating gut microbiota[J].Environmental Pollution, 2022, 292(Pt B): 118477. DOI:10.1016/j.envpol.2021.118477.
[63] WANG K D, ZHANG C H, ZHANG B F, et al. Gut dysfunction may be the source of pathological aggregation of alpha-synuclein in the central nervous system through Paraquat exposure in mice[J].Ecotoxicology and Environmental Safety, 2022, 246: 114152.DOI:10.1016/j.ecoenv.2022.114152.
[64] DEL CASTILO I, NEUMANN A S, LEMOS F S, et al. Lifelong exposure to a low-dose of the glyphosate-based herbicide RoundUp®causes intestinal damage, gut dysbiosis, and behavioral changes in mice[J]. International Journal of Molecular Sciences, 2022, 23(10):5583. DOI:10.3390/ijms23105583.
[65] YUE Y, ZHANG H D, DENG P, et al. Environmental cadmium exposure facilitates mammary tumorigenesis via reprogramming gut microbiota-mediated glutamine metabolism in MMTV-Erbb2 mice[J].Science of the Total Environment, 2023, 897: 165348. DOI:10.1016/j.scitotenv.2023.165348.
[66] YANG J S, CHEN W, SUN Y, et al. Effects of cadmium on organ function, gut microbiota and its metabolomics profile in adolescent rats[J]. Ecotoxicology and Environmental Safety, 2021, 222: 112501.DOI:10.1016/j.ecoenv.2021.112501.
[67] MA Z, GAO X, YANG X, et al. Low-dose florfenicol and copper combined exposure during early life induced health risks by affecting gut microbiota and metabolome in SD rats[J]. Ecotoxicology and Environmental Safety, 2022, 245: 114120. DOI:10.1016/j.ecoenv.2022.114120.
[68] ZHANG S, SHEN Y, WANG S N, et al. Responses of the gut microbiota to environmental heavy metal pollution in tree sparrow(Passer montanus) nestlings[J]. Ecotoxicology and Environmental Safety, 2023, 264: 115480. DOI:10.1016/j.ecoenv.2023.115480.
[69] WU N, WANG X B, XU X Y, et al. Effects of heavy metals on the bioaccumulation, excretion and gut microbiome of black soldier fly larvae (Hermetia illucens)[J]. Ecotoxicology and Environmental Safety, 2020, 192: 110323. DOI:10.1016/j.ecoenv.2020.110323.
[70] XU Y J, YU Y P, ZHOU Q, et al. Disturbance of gut microbiota aggravates cadmium-induced neurotoxicity in zebrafish larvae through V-ATPase[J]. Science of the Total Environment, 2023, 891: 164074.DOI:10.1016/j.scitotenv.2023.164074.
[71] THOMAS J C IV, KIERAN T J, FINGER J W Jr, et al. Unveiling the gut microbiota and resistome of wild cotton mice, Peromyscus gossypinus, from heavy metal- and radionuclide-contaminated sites in the southeastern United States[J]. Microbiology Spectrum, 2021, 9(1):e0009721. DOI:10.1128/Spectrum.00097-21.
[72] YU J, CHEN L, WU B. Size-specific effects of microplastics and lead on zebrafish[J]. Chemosphere, 2023, 337: 139383. DOI:10.1016/j.chemosphere.2023.139383.
[73] LI A Y, AN Z G, LI C Y, et al. Salt-contaminated water exposure induces gut microbial dysbiosis in chickens[J]. Ecotoxicology and Environmental Safety, 2023, 254: 114731. DOI:10.1016/j.ecoenv.2023.114731.
[74] LI Y T, ZHANG R X, LI X, et al. Exposure to the environmental pollutant ammonia causes changes in gut microbiota and inflammatory markers in fattening pigs[J]. Ecotoxicology and Environmental Safety,2021, 208: 111564. DOI:10.1016/j.ecoenv.2020.111564.
[75] QI M, WANG N, XIAO Y X, et al. Ellagic acid ameliorates paraquatinduced liver injury associated with improved gut microbial profile[J].Environmental Pollution, 2022, 293: 118572. DOI:10.1016/j.envpol.2021.118572.
[76] LAUE H E, BRENNAN K J M, GILLET V, et al. Associations of prenatal exposure to polybrominated diphenyl ethers and polychlorinated biphenyls with long-term gut microbiome structure: a pilot study[J]. Environmental Epidemiology, 2019, 3(1): e039. DOI:10.1097/EE9.0000000000000039.
[77] EGGERS S, SAFDAR N, SETHI A K, et al. Urinary lead concentration and composition of the adult gut microbiota in a crosssectional population-based sample[J]. Environment International,2019, 133: 105122. DOI:10.1016/j.envint.2019.105122.
[78] CHASSAING B, COMPHER C, BONHOMME B, et al. Randomized controlled-feeding study of dietary emulsifier carboxymethylcellulose reveals detrimental impacts on the gut microbiota and metabolome[J].Gastroenterology, 2022, 162(3): 743-756. DOI:10.1053/j.gastro.2021.11.006.
[79] LAI H, LI Y F, HE Y F, et al. Effects of dietary fibers or probiotics on functional constipation symptoms and roles of gut microbiota:a double-blinded randomized placebo trial[J]. Gut Microbes, 2023,15(1): 2197837. DOI:10.1080/19490976.2023.2197837.
[80] ANYANSI C, STRAUB T J, MANSON A L, et al. Computational methods for strain-level microbial detection in colony and metagenome sequencing data[J]. Frontiers in Microbiology, 2020, 11: 1925.DOI:10.3389/fmicb.2020.01925.
[81] ZHANG Y C, THOMPSON K N, HUTTENHOWER C, et al.Statistical approaches for differential expression analysis in metatranscriptomics[J]. Bioinformatics, 2021, 37(Suppl 1): i34-i41.DOI:10.1093/bioinformatics/btab327.
[82] FARAG M A, BAKY M H, VON BERGEN M, et al. The use of omics in monitoring food gut microbiota interaction outcomes: a review of novel trends and technologies[J]. Current Opinion in Food Science,2023, 52: 101064. DOI:10.1016/j.cofs.2023.101064.
[83] SUN J C, FANG R C, WANG H, et al. A review of environmental metabolism disrupting chemicals and effect biomarkers associating disease risks: where exposomics meets metabolomics[J]. Environment International, 2022, 158: 106941. DOI:10.1016/j.envint.2021.106941.
[84] GAO B, CHI L, MAHBUB R, et al. Multi-omics reveals that lead exposure disturbs gut microbiome development, key metabolites, and metabolic pathways[J]. Chemical Research in Toxicology, 2017, 30(4):996-1005. DOI:10.1021/acs.chemrestox.6b00401.
[85] ZENG X, ZENG Z J, WANG Q H, et al. Alterations of the gut microbiota and metabolomics in children with e-waste lead exposure[J]. Journal of Hazardous Materials, 2022, 434: 128842.DOI:10.1016/j.jhazmat.2022.128842.
[86] WU C H, FANG F, YU Y, et al. Multi-omics analyses of serum metabolome, urine metabolome and gut microbiome reveal dysregulated glycerophospholipid metabolism in subacute cadmiumexposed wistar rats[J]. Toxicology, 2023, 494: 153594. DOI:10.1016/j.tox.2023.153594.
[87] ZHANG L, ZHOU Y T, SONG Z W, et al. Mercury induced tissue damage, redox metabolism, ion transport, apoptosis, and intestinal microbiota change in red swamp crayfish (Procambarus clarkii):application of multi-omics analysis in risk assessment of Hg[J].Antioxidants, 2022, 11(10): 1944. DOI:10.3390/antiox11101944.
[88] MESNAGE R, TEIXEIRA M, MANDRIOLI D, et al. Use of shotgun metagenomics and metabolomics to evaluate the impact of glyphosate or roundup MON 52276 on the gut microbiota and serum metabolome of sprague-dawley rats[J]. Environmental Health Perspectives, 2021,129(1): 17005. DOI:10.1289/EHP6990.
[89] DEFOIS C, RATEL J, GARRAIT G, et al. Food chemicals disrupt human gut microbiota activity and impact intestinal homeostasis as revealed by in vitro systems[J]. Scientific Reports, 2018, 8(1): 11006.DOI:10.1038/s41598-018-29376-9.
[90] DALIRI E B M, OFOSU F K, CHELLIAH R, et al. Challenges and perspective in integrated multi-omics in gut microbiota studies[J].Biomolecules, 2021, 11(2): 300. DOI:10.3390/biom11020300.
[91] SHI Y, WANG G P, LAU H C H, et al. Metagenomic sequencing for microbial DNA in human samples: emerging technological advances[J]. International Journal of Molecular Sciences, 2022, 23(4):2181. DOI:10.3390/ijms23042181.
[92] FASSARELLA M, BLAAK E E, PENDERS J, et al. Gut microbiome stability and resilience: elucidating the response to perturbations in order to modulate gut health[J]. Gut, 2021, 70(3): 595-605.DOI:10.1136/gutjnl-2020-321747.
[93] HASAN N, YANG H Y. Factors affecting the composition of the gut microbiota, and its modulation[J]. PeerJ, 2019, 7: e7502. DOI:10.7717/peerj.7502.
[94] LINDELL A E, ZIMMERMANN-KOGADEEVA M, PATIL K R.Multimodal interactions of drugs, natural compounds and pollutants with the gut microbiota[J]. Nature Reviews Microbiology, 2022, 20(7):431-443. DOI:10.1038/s41579-022-00681-5.
[95] VUJKOVIC-CVIJIN I, SKLAR J, JIANG L J, et al. Host variables confound gut microbiota studies of human disease[J]. Nature, 2020,587: 448-454. DOI:10.1038/s41586-020-2881-9.
[96] SARIN S K, PANDE A, SCHNABL B. Microbiome as a therapeutic target in alcohol-related liver disease[J]. Journal of Hepatology, 2019,70(2): 260-272. DOI:10.1016/j.jhep.2018.10.019.
[97] MAITRE L, BUSTAMANTE M, HERNÁNDEZ-FERRER C, et al.Multi-omics signatures of the human early life exposome[J]. Nature Communications, 2022, 13(1): 7024. DOI:10.1038/s41467-022-34422-2.
[98] ZHANG Q L, DU X H, LI H C, et al. Cardiovascular effects of trafficrelated air pollution: a multi-omics analysis from a randomized,crossover trial[J]. Journal of Hazardous Materials, 2022, 435: 129031.DOI:10.1016/j.jhazmat.2022.129031.
[99] FAN L J, XIA Y Y, WANG Y X, et al. Gut microbiota bridges dietary nutrients and host immunity[J]. Science China Life Sciences, 2023,66(11): 2466-2514. DOI:10.1007/s11427-023-2346-1.
[100] QU R Z, ZHANG Y, MA Y P, et al. Role of the gut microbiota and its metabolites in tumorigenesis or development of colorectal cancer[J]. Advanced Science, 2023, 10(23): e2205563. DOI:10.1002/advs.202205563.
[101] WONG C C, YU J. Gut microbiota in colorectal cancer development and therapy[J]. Nature Reviews Clinical Oncology, 2023, 20(7): 429-452. DOI:10.1038/s41571-023-00766-x.
[102] BUNDGAARD-NIELSEN C, HAGSTRØM S, SØRENSEN S.Interpersonal variations in gut microbiota profiles supersedes the effects of differing fecal storage conditions[J]. Scientific Reports,2018, 8(1): 17367. DOI:10.1038/s41598-018-35843-0.
[103] KAZANTSEVA J, MALV E, KALEDA A, et al. Optimisation of sample storage and DNA extraction for human gut microbiota studies[J]. BMC Microbiology, 2021, 21(1): 158. DOI:10.1186/s12866-021-02233-y.
[104] GONG J H, SHEN Y, ZHANG H C, et al. Gut microbiota characteristics of people with obesity by meta-analysis of existing datasets[J]. Nutrients, 2022, 14(14): 2993. DOI:10.3390/nu14142993.
[105] JIN H, YOU L J, ZHAO F Y, et al. Hybrid, ultra-deep metagenomic sequencing enables genomic and functional characterization of lowabundance species in the human gut microbiome[J]. Gut Microbes,2022, 14(1): 2021790. DOI:10.1080/19490976.2021.2021790.
[106] GAO B, CHI L, ZHU Y X, et al. An introduction to next generation sequencing bioinformatic analysis in gut microbiome studies[J].Biomolecules, 2021, 11(4): 530. DOI:10.3390/biom11040530.
[107] AMOS G C A, LOGAN A, ANWAR S, et al. Developing standards for the microbiome field[J]. Microbiome, 2020, 8(1): 98. DOI:10.1186/s40168-020-00856-3.
[108] KNIGHT R, VRBANAC A, TAYLOR B C, et al. Best practices for analysing microbiomes[J]. Nature Reviews Microbiology, 2018, 16(7):410-422. DOI:10.1038/s41579-018-0029-9.
[109] COSTEA P I, ZELLER G, SUNAGAWA S, et al. Towards standards for human fecal sample processing in metagenomic studies[J]. Nature Biotechnology, 2017, 35(11): 1069-1076. DOI:10.1038/nbt.3960.
[110] TOURLOUSSE D M, NARITA K, MIURA T, et al. Validation and standardization of DNA extraction and library construction methods for metagenomics-based human fecal microbiome measurements[J].Microbiome, 2021, 9(1): 95. DOI:10.1186/s40168-021-01048-3.
[111] LYNCH S V, NG S C, SHANAHAN F, et al. Translating the gut microbiome: ready for the clinic?[J]. Nature Reviews Gastroenterology & Hepatology, 2019, 16(11): 656-661. DOI:10.1038/s41575-019-0204-0.
